goal: results of preliminary look at UCSD HIV-ENV sequencing
- goal: examine ENV data from UCSD using clustering consensus for CROI abstract due Oct 8th. - first look at the single patient with three time points, P018 months 3, 22 and 33 to get initial result that might be included in CROI abstract. - The expected result: "the one patient for which we have longitudinal data is monoinfected, and has a strong autologous antibody response: we "expect" to see greater numbers of supported variants as time increases." - Summary: - The complexity of the sample mixtures appears to increase over time. - The subpopulation of larger-delete variants appears to increase and change over time. Next ================================Inputs and Methods
- I used the HXB2 reference (base 0 to 9718). After alignments, I trimmed the reference to the covered bases 5960:9160. Here is the trimmed reference: hiv_hxb2_ENV.fasta - I examined these three runs: P018_3m_5pM, P018_22m_10pM, P018_33m_5pM. - ClusteringConsensus was used to align reads to the trimmed reference and only use CCS reads that fully-spanned 3169 or more of the 3201 total reference bases (I allowed a 1% loss on the ends). Multiple alignments are generated and then reads in the alignments are clustered to look for structure. Various statistics are reported. Next ================================Mapping statistics
- Number of CCS reads that completely cover the 3.2kb of the reference missing at most 1% on the ends:
Number Sample
7091 clucon_HBX2ref_P018_3m
7896 clucon_HBX2ref_P018_22m
8624 clucon_HBX2ref_P018_33m
- A good number of full-length CCS hits: 7091 to 8624 coverage per
chip.
- Here are the simple stats for those hits:
clucon_HBX2ref_P018_3m/alignments.filterFull
clucon_HBX2ref_P018_22m/alignments.filterFull
clucon_HBX2ref_P018_33m/alignments.filterFull
Next
================================
Variant Positions
- Number of positions that are likely to contain minor variants according to simple entropy threshold:
Num Sample
93 clucon_HBX2ref_P018_3m
135 clucon_HBX2ref_P018_22m
176 clucon_HBX2ref_P018_33m
- The number of variant positions increases
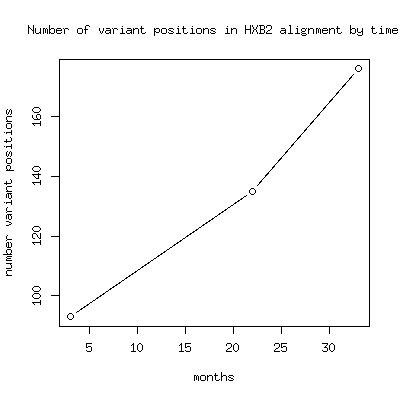 - Here are the list of variant multiple alignment positions by this
rough entropy measure:
clucon_HBX2ref_P018_3m/distjob.usecols
clucon_HBX2ref_P018_22m/distjob.usecols
clucon_HBX2ref_P018_33m/distjob.usecols
Note that (reference position 1 based)=(alignment position)/5 because
I gap out inserts with 4 columns.
Next
================================
- Here are the list of variant multiple alignment positions by this
rough entropy measure:
clucon_HBX2ref_P018_3m/distjob.usecols
clucon_HBX2ref_P018_22m/distjob.usecols
clucon_HBX2ref_P018_33m/distjob.usecols
Note that (reference position 1 based)=(alignment position)/5 because
I gap out inserts with 4 columns.
Next
================================
Clustering
- Examine the complete-linkage clustering of all reads on these variant positions (3m, 22m, 33m):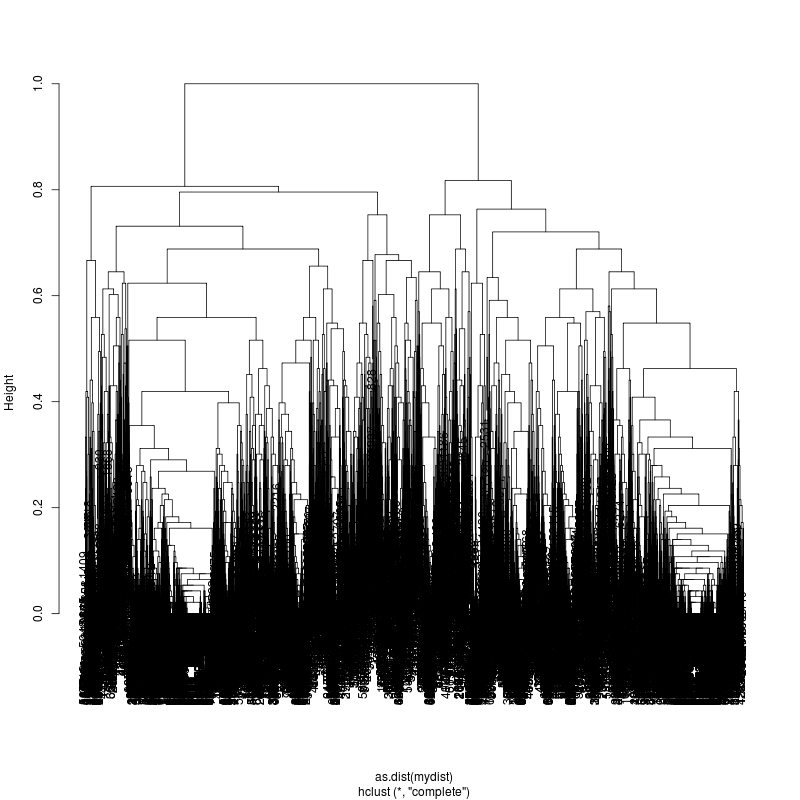
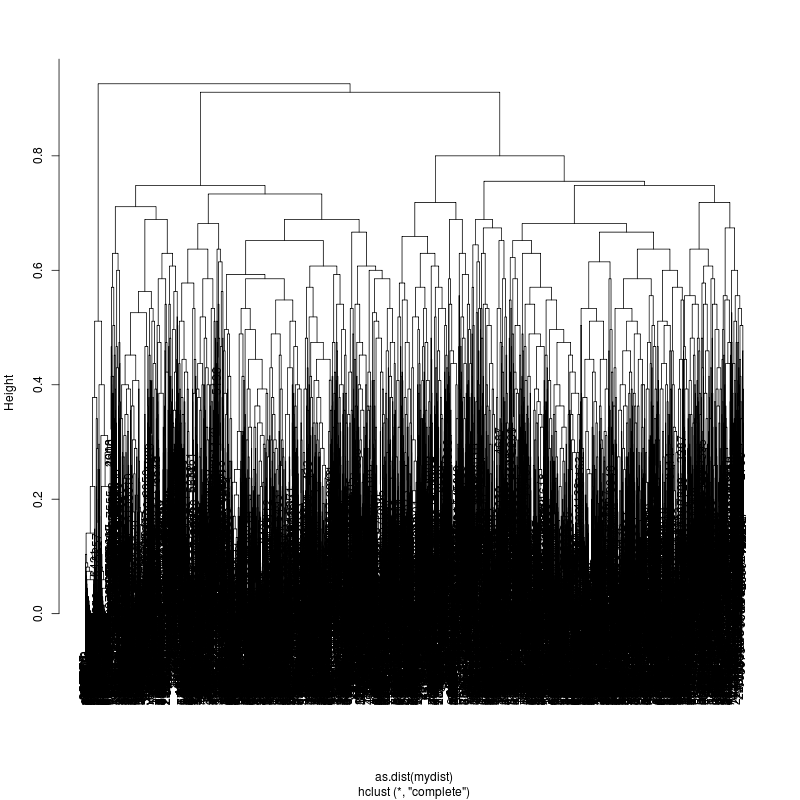
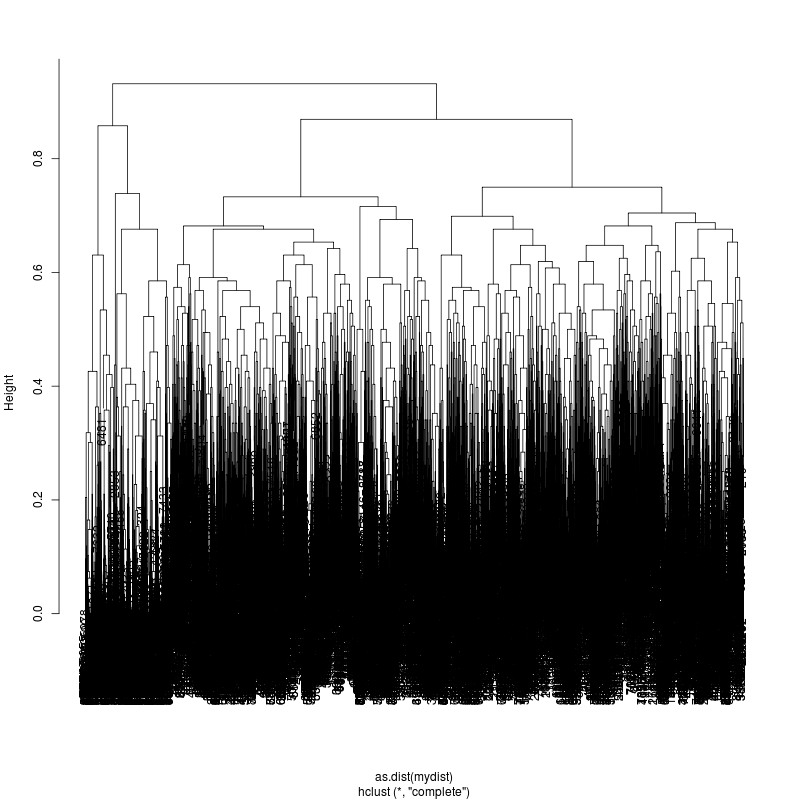 - Each column is a full-length amplicon-spanning read and the y-axis
represents the distance which is the fraction of variant positions
that disagree (0=identical, 1=completely different over the 93,135, or
176 variant positions). For example, a join distance of 0.8 between
subclusters says that every pairwise distance in the subtree is less
than 0.8.
- The initial 3m sample is fairly complex. The complexity appears to
be roughly increasing. More work is needed to stratify clusters using
bionomial noise bounds and call more precise subspecies.
Next
================================
- Each column is a full-length amplicon-spanning read and the y-axis
represents the distance which is the fraction of variant positions
that disagree (0=identical, 1=completely different over the 93,135, or
176 variant positions). For example, a join distance of 0.8 between
subclusters says that every pairwise distance in the subtree is less
than 0.8.
- The initial 3m sample is fairly complex. The complexity appears to
be roughly increasing. More work is needed to stratify clusters using
bionomial noise bounds and call more precise subspecies.
Next
================================
Large Deletions
- How many reads have more than 10% deleted positions with respect to hxb2?
Sample FractionHXB2Deleted
3m 0.05330701
22m 0.05889058
33m 0.1621058
33m has a triple the fraction of reads that have deletions of more
than 10% HXB2 positions.
- Fraction of reads that delete at each position in the reference:
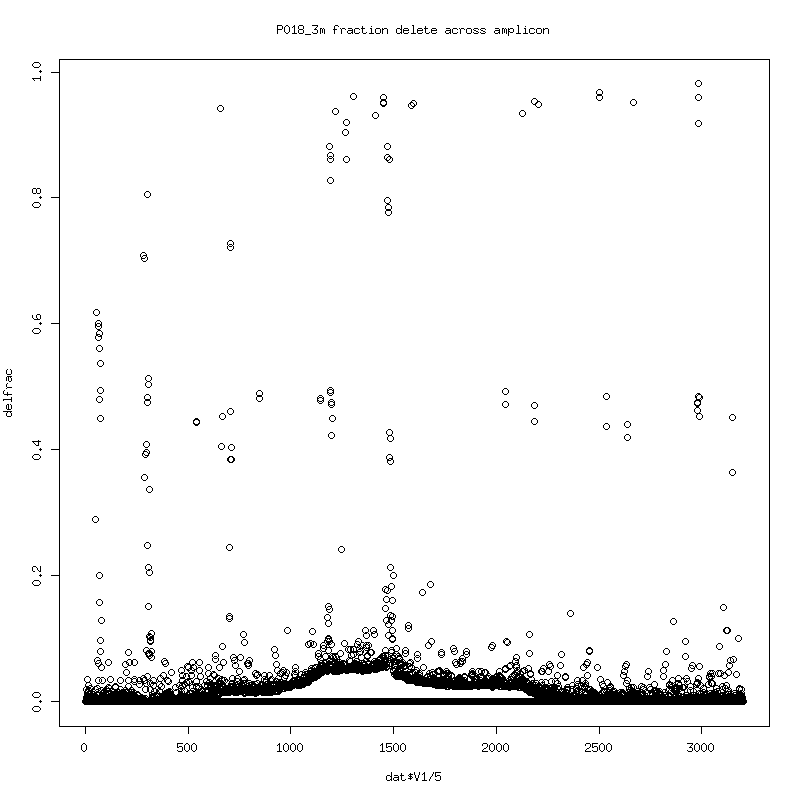
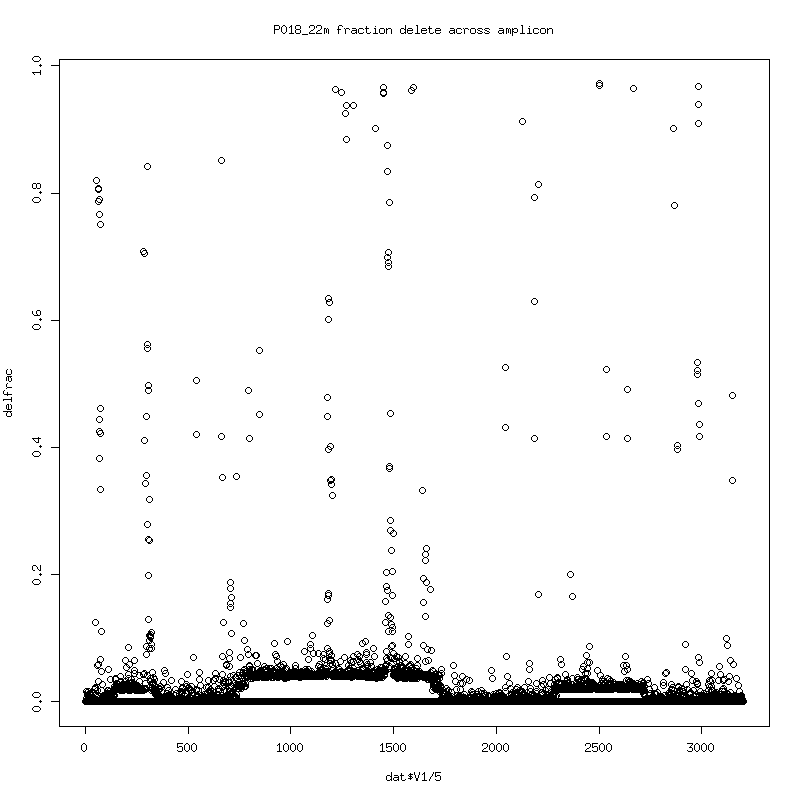
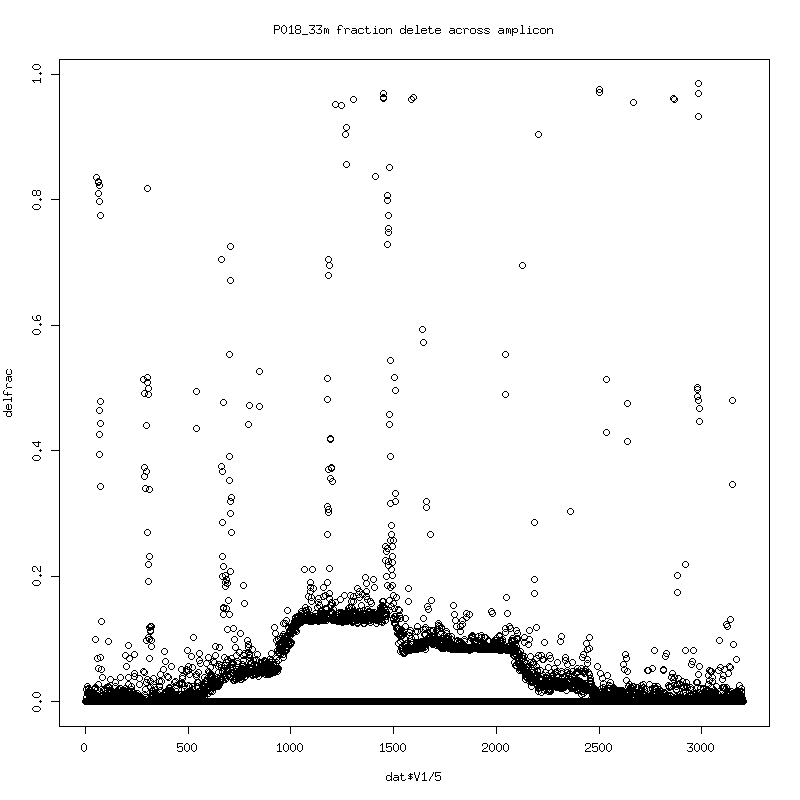 There seems to be an evolution of delete variants (consistent deletes
happens for many 100s of bases...) The consistent spikes might be
somewhat consistent differences with the HXB2 reference and local
alignment artifacts.
Next
================================
There seems to be an evolution of delete variants (consistent deletes
happens for many 100s of bases...) The consistent spikes might be
somewhat consistent differences with the HXB2 reference and local
alignment artifacts.
Next
================================
Alignment Entropy
- Plot of the entropy of each column in the alignment. (Higher entropy means more variability in observed bases. 0 entropy would indicate that a single base identity was observed with nothing else).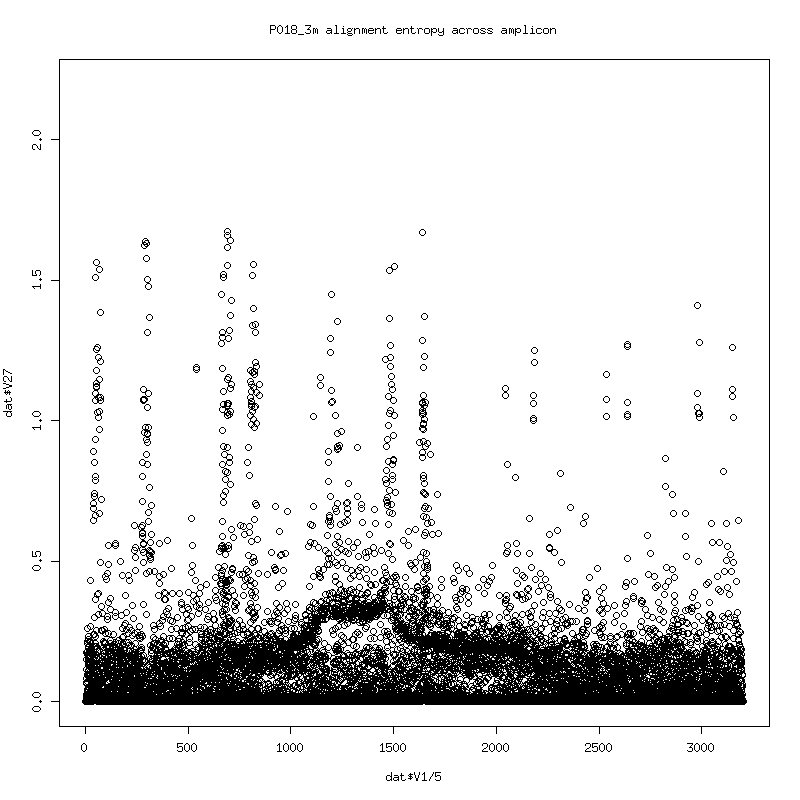
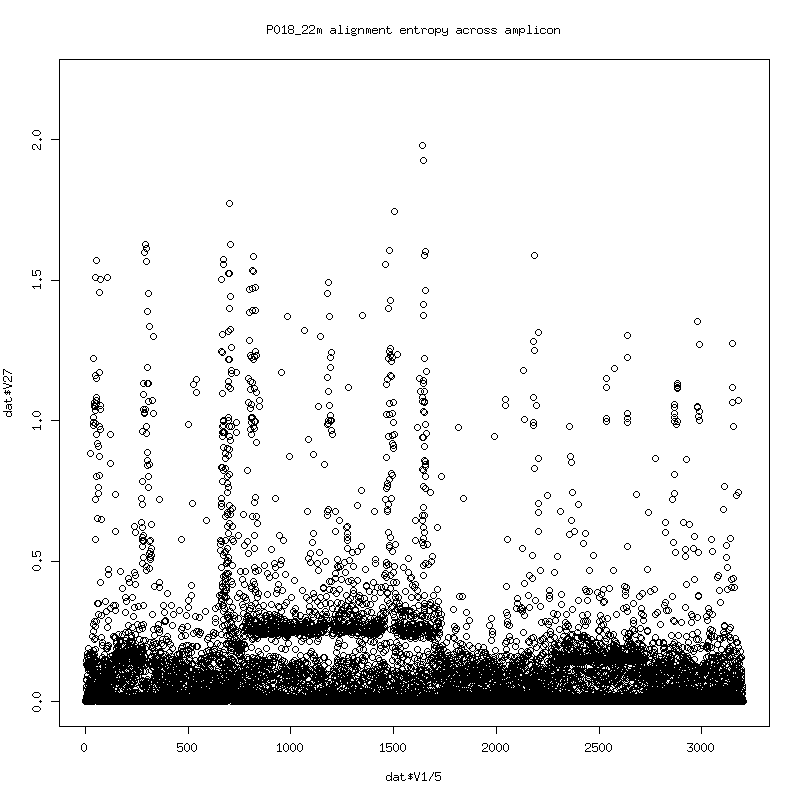
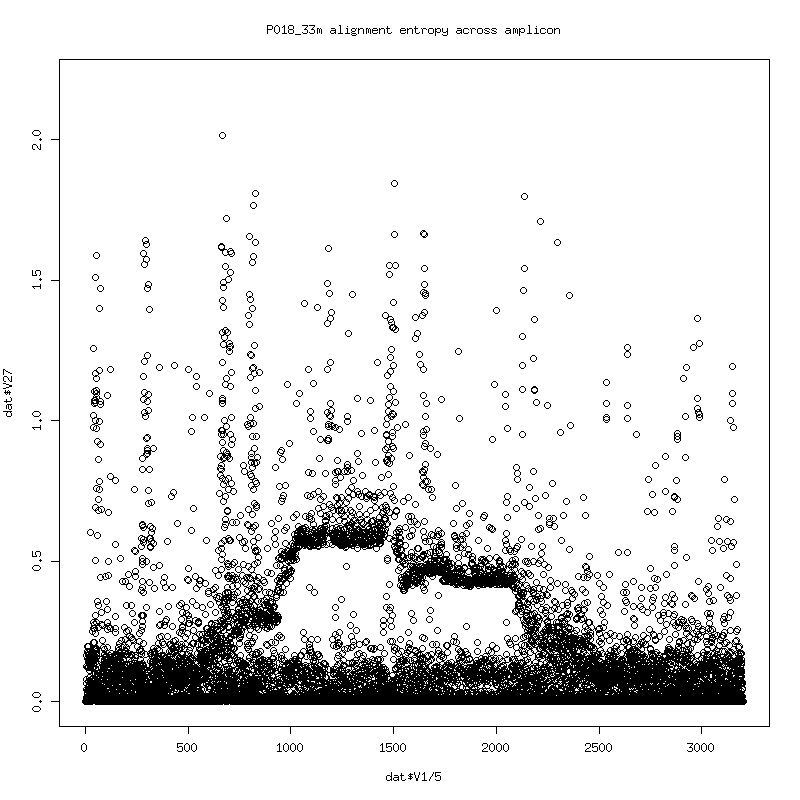 - The entropy somewhat follows the deletes. The consistent spikes
might be somewhat consistent differences with the HXB2 reference and
local alignment artifacts.
End
- The entropy somewhat follows the deletes. The consistent spikes
might be somewhat consistent differences with the HXB2 reference and
local alignment artifacts.
End