This page shows mutual information estimates between every base of the HIV genome (9032 bases in these experiments). The image is zoomable down to single base resolution with an interface much like Google Maps.
The information in this visualization is based on a single sample from a single patient where we estimate there to be about six major HIV genome-types in the sample.
PacBio long reads allow this estimate of mutual information because we sequence almost the entire HIV genome from single molecules in single reads. The artifacts at the lower-left and upper-right are caused by small sample where we don't capture to exactly the ends of the genome (caused by fewer reads at the longest lengths as well as sample prep issues).
Mutual information measures the information above independence assumptions. This might be caused by selection pressures (structural, coding, ..) as HIV explores the mutation landscape.
Clicking will show what genes are annotated at those genome positions in the text to the right of the image. geneBhit represents the x-axis while geneEhit represents the y-axis.
The javascript image tiling code is taken from GSV 1.0: http://mike.teczno.com/giant/pan/. I have had better luck using Chrome and have seen issues in Firefox.
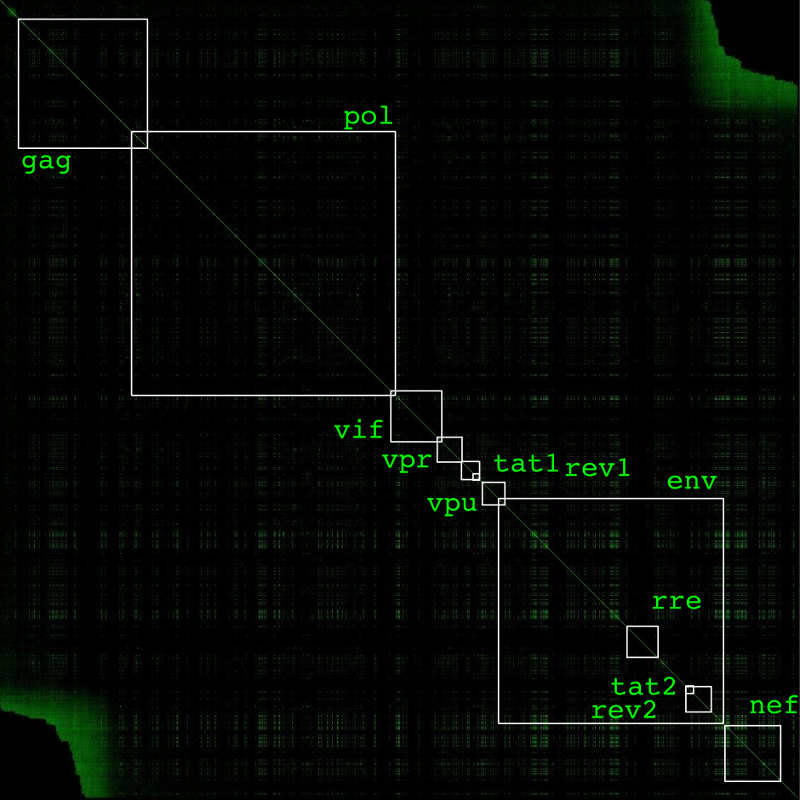
Here is a image annotated with a subset of the gene locations: